This is Project 2 in our overall research scheme.
This Project studies somatic cell evolution in distinct, small human replicative units (intestinal crypts and tumor glands). The compartmentalization of cells into small replicative units can modify evolution because selection and drift (random cell turnover) is limited to immediately adjacent cells. The advantages of analyzing small replicative units are that experimentally they large enough to measure with conventional methods yet small enough to simulate in detail. Characterizing somatic cell evolution in replicative units can lead to better understanding of tumor evolution because selection or drift first occurs at the cell-cell level.
We are sampling multiple colon crypts from differently aged individuals and multiple tumor glands from opposite sides of 20 colorectal tumors (adenomas and CRCs). Crypts from different species will also be compared with the human data. For each crypt or gland we will document genetic alterations (whole genome sequencing), epigenetic alterations (ATAC-seq), and expression. Combining epigenetic and genetic evolution is important because phenotypic plasticity can mimic clonal evolution when epigenetic modulation allows a single genome to express multiple phenotypes. This data will be modeled through simulations to better understand somatic cell proliferation (division and death) within small replicative units, specifically to determine whether selection or neutral drift are more common during normal human aging and tumor progression.
This Project is also comparing the evolutionary biology of fibroblasts. We are testing the predictions of Project 1 for the mutation rates, proliferation rates, apoptosis rates, DNA damage response, and stress responses of early passage, primary fibroblasts from species with a wide variety of life history traits. In addition, we are testing for functional differences in cancer associated pathways that have evidence of selection in specific species identified by Project 1.
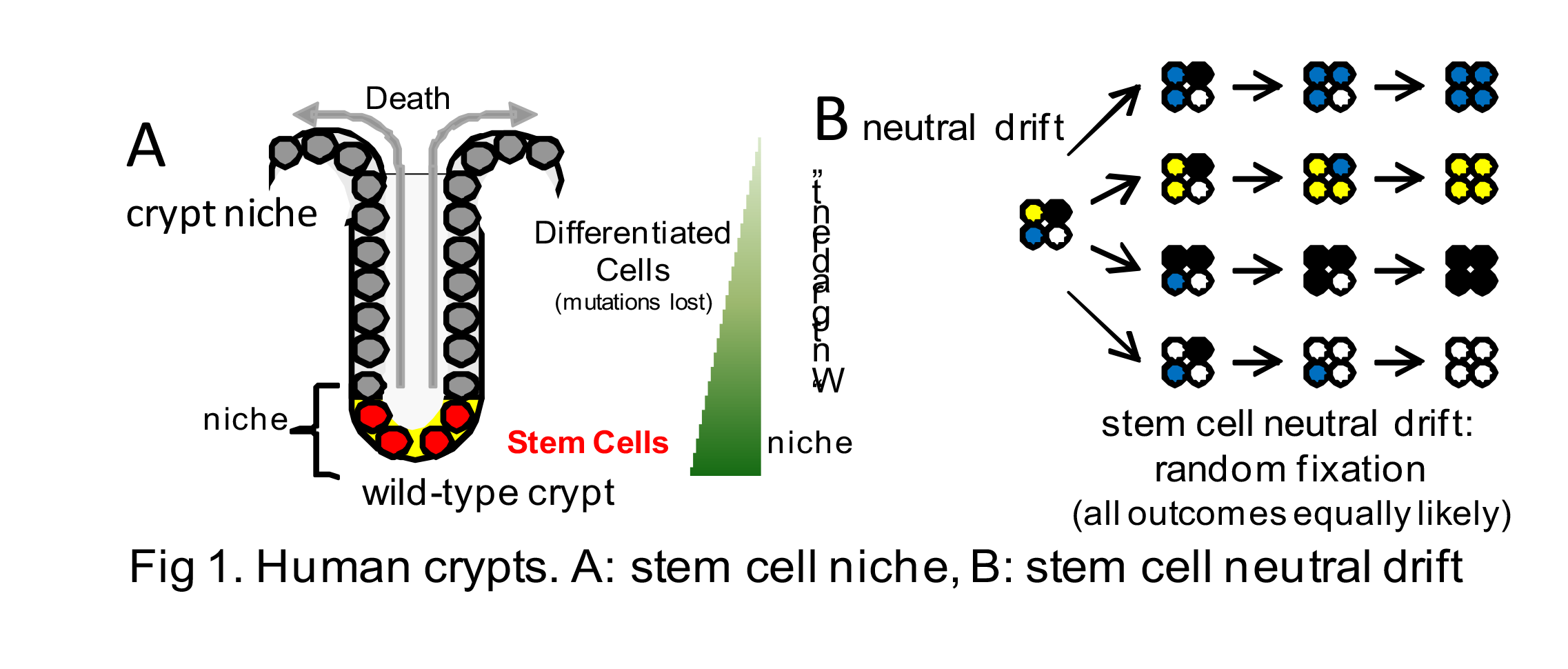
Somatic Cell Evolution in Small Human Replicative Units
Investigators:
Darryl Shibata (Project Leader and Co-PI)
Joshua Schiffman (Project Leader)
Trevor Graham (Project Leader)
Lisa Abegglen