This is Project 3 in our overall scheme.
Cancer is an evolutionary process where a single cell grows into a visible tumor after it has acquired multiple driver alterations. In this Project, we are developing predictive biomarkers for colorectal cancer (CRC) outcomes. The innovation is that the biomarkers will be direct measures of evolution. Such evolution-based biomarkers also provide mechanistic understanding for what aspects of evolution most impact survival. To fully characterize tumor evolution, it is necessary to measure both how tumor cells evolve and the host ecology.
In Aim 1, mutations detected by whole exome sequencing of CRC cohorts with long-term follow-up are being classified as public (clonal) and private (subclonal). CRCs are then subclassified with a newly developed algorithm that infers selection (positive, neutral, negative) based on private mutation frequencies, where selection preferentially increases (positive) or decreases (negative) subclone subsets. In Aim 2, we are scanning microscope sections from the above tumors, using a unique automated platform that can identify cells and quantify tumor microenvironments with respect to lymphocytes and stromal cells. To determine if host ecological heterogeneity reflects responses to specific tumor subclones, we are overlaying private mutation distributions on the same microscope slides.
As the project progresses, we will combine tumor evolution and the host reaction into a single evolution-ecology (Evo-Eco) index that summarizes the underlying evolutionary struggle. For example, patients with aggressive tumors (positive selection) and supportive environments are likely to have poorer outcomes relative to patients with tumors under negative selection and repressive environments. We will validate our Evo-Eco index on another cohort of CRCs. If successful, these studies will yield improved CRC predictive biomarkers.
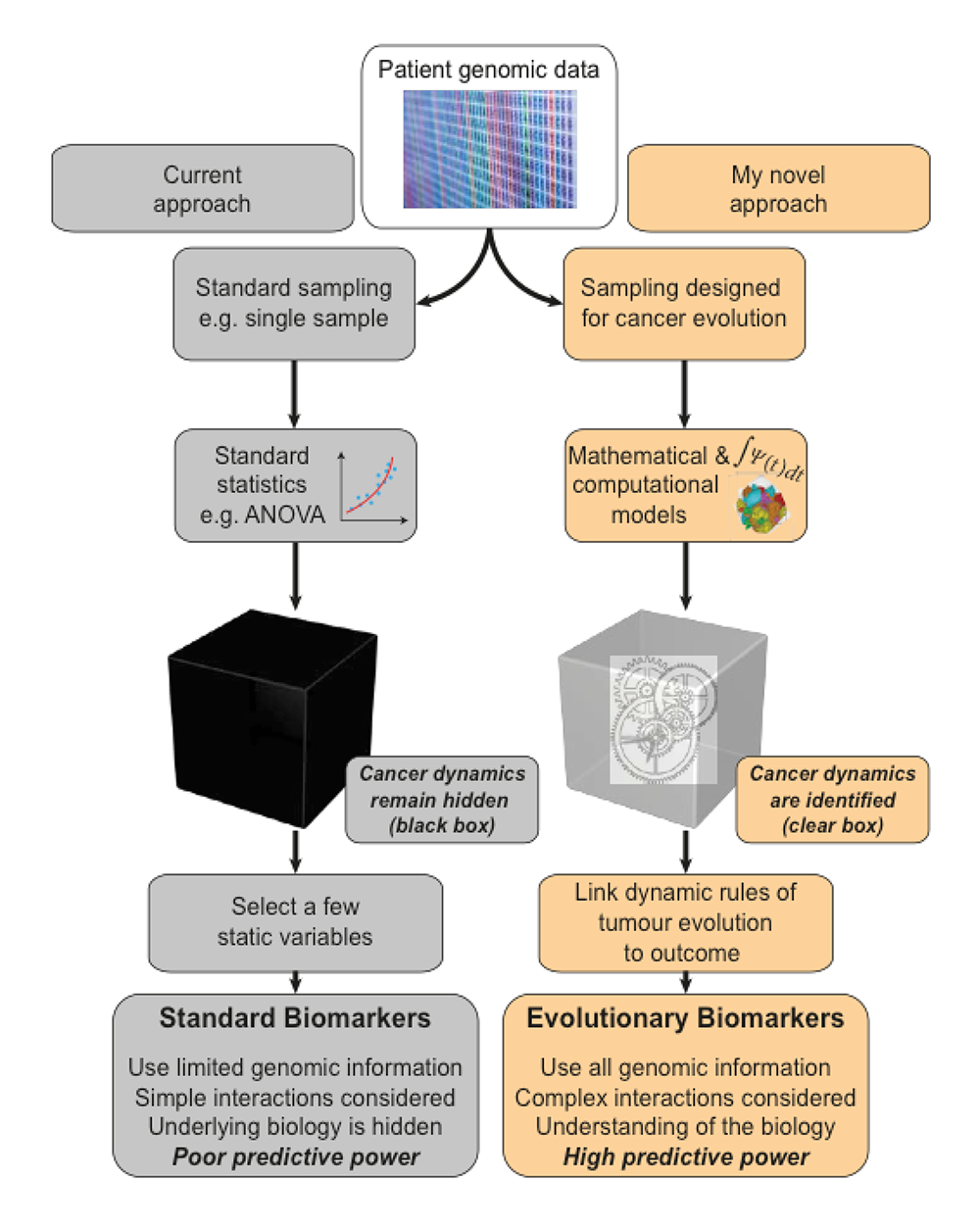
Neoplastic Cell Evolution
Investigators
Trevor Graham (Project Leader)
Yinyin Yuan (Project Leader)
Darryl Shibata (Project Leader and Co-PI)
Andrea Sottoriva
Li Liu
Christina Curtis